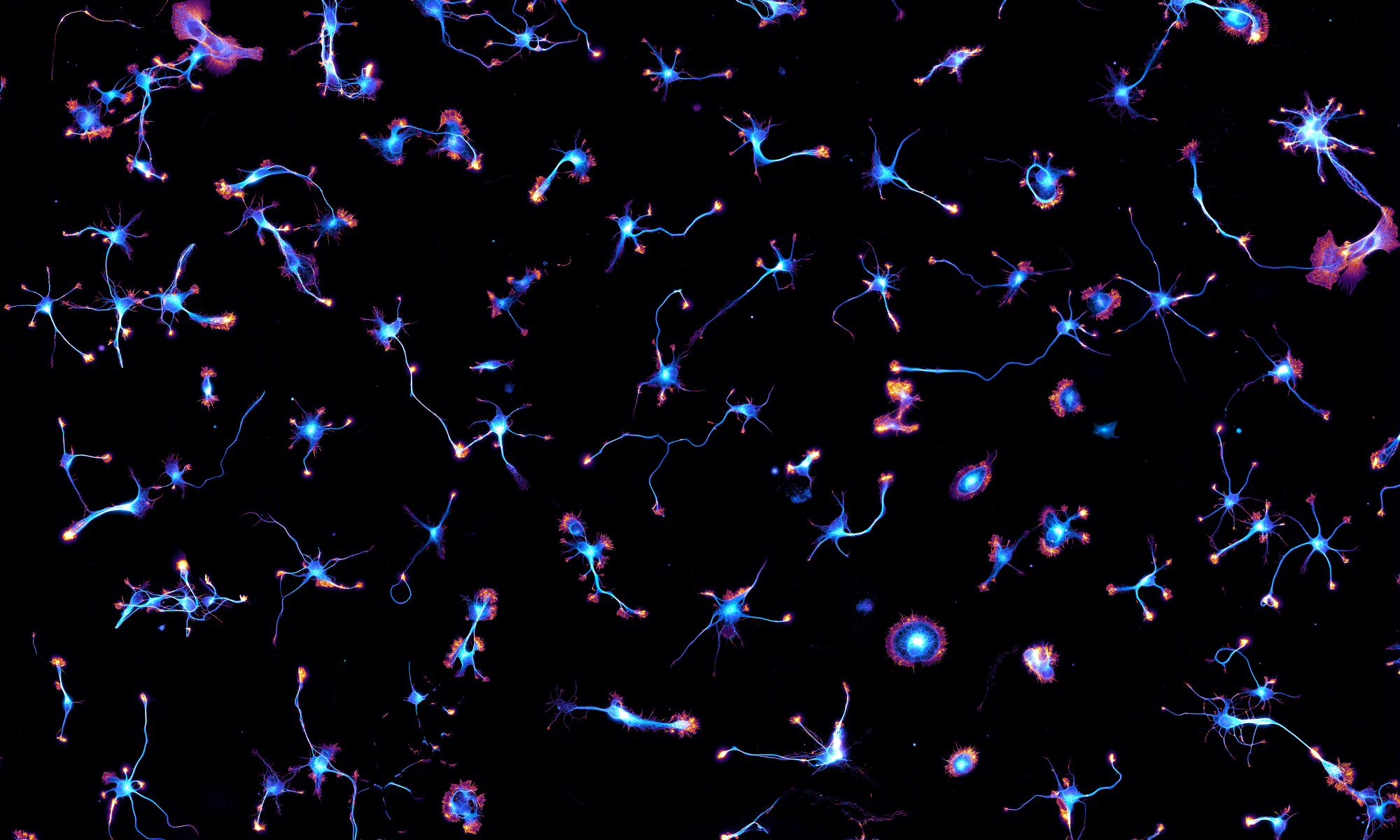
Our latest preprint is out, a collaboration with our long-standing collaborator Subhojit Roy and his lab, in particular the talented postdoc Archan Ganguly. We uncover how clathrin is transported along axons as assembled structures that are unrelated to endocytosis: the “transport packets”. In this work, we pushed DNA-PAINT to image these axonal clathrin packets. Here they are by EM of APEX-clathrin on top and by DNA-PAINT of endogenous clathrin at the bottom. We also repurposed the ChimeraX software to render PAINT data and see inside axons!

Where are the packets going? Inside presynapses! We visualized clathrin in presynapses – 3D DNA-PAINT could clearly separate presynaptic and postsynaptic clusters. With ChimeraX we rendered and measured these presynaptic clathrin packets – similar in size to the ones along axons.

Check the preprint for the whole story – there’s so much more in there. Great work from Florian and Ghislaine in the team, congrats everyone!
Ganguly A, Wernert F, Phan S, Boassa D, Das U, Sharma R, Caillol G, Han X, Yates JR, Ellisman M, Leterrier C, Roy S.
Mechanistic Determinants of Slow Axonal Transport and Presynaptic Targeting of Clathrin Packets.
bioRxiv, 2020 Feb 20.
doi: 10.1101/2020.02.20.958140